 Projects: Organs - Alzheimer's Disease
Projects: Organs - Alzheimer's Disease
Alzheimer's Disease markers from structural MRI

Mads Nielsen, PhD
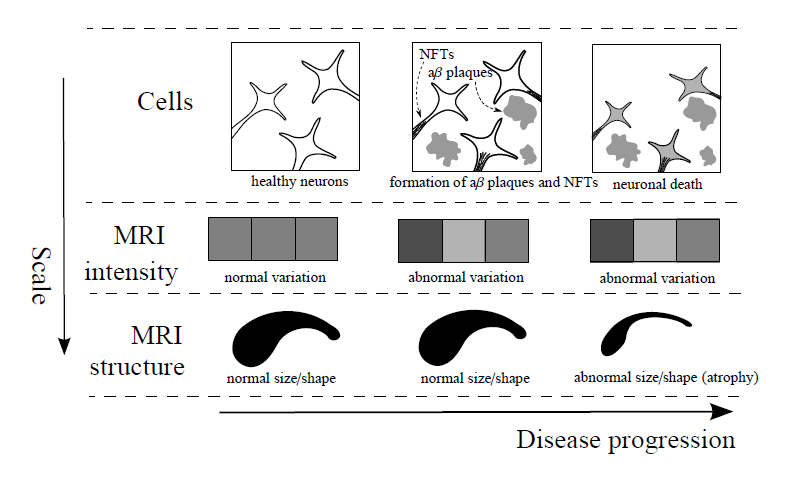
In dementia trials cognitive decline as end point has recently been complimented or even substituted by MRI-based brain atrophy measurements of the whole brain or specific brain areas such as the hippocampus severely affected by AD.
- The Atrophiq framework by Biomediq offers computer-based quantification of structural (T1 weighted) brain MRI:
- Atrophy computation based on bias correction, skull stripping, and segmentation of whole brain and hippocampus according to the Harmonized Hippocampus Protocol (HHP). Accuracy and precision are surpassing automated and radiologist's results in the literature. Symmetric longitudinal atrophy measurements are based on proprietary and patented technology.
- Atrophy markers include volumetry and atrophy of whole brain, hippocampus, the medial temporal lope, ventricles, and several other brain structures.
- The range of markers covers prognostic markers for study population selection, efficacy markers for treatment evaluation, and exploratory markers for investigation of treatment mode of action.
- MRI-based analysis of hippocampal structure and shape, and quantification of neuro-vascular pathologies. Experimental markers include glucose and amyloid PET, confined to cortical areas defined by the structural MRI.
- Efficient workflow since the radiologist reading bottleneck is replaced by automation. If needed, all scans from large phase III trials can be quantified within a week at the Biomediq high performance computing facility.
Validation and Technical Performance
The Atrophiq framework has been validated on several population-based studies including the Alzheimer's Disease Neuroimaging Initiative (ADNI), the The Australian Imaging, Biomarker & Lifestyle Flagship Study of Ageing (AIBL) and the The Metropolit 1953 Danish Male Birth Cohort. Results have been presented at leading conferences like AAIC and RSNA.
Hippocampus Segmentation Accuracy
In the literature manual inter-rater dice volume overlap is reported to be 80-90% for repeated segmentations of the hippocampus by expert raters. The Atrophiq produces overlap of 87% with manual segmentations produced by the Harmonized Hippocampal Protocol expert group. This compares to the de facto standard of FreeSurfer with a significantly lower overlap of 78%.
Efficacy Markers
A standard sub-set of the ADNI trial consists of 504 subjects diagnosed as 169 normal controls (NC), 234 mild cognitively impaired (MCI), and 101 with AD diagnosis. They were followed for 2 years with annual MRI. A model trial with 25% drug efficacy is used for simulating power analysis. Atrophiq improves the signal to noise ratio of hippocampal atrophy. Sample sizes (SS) are reduced compared to Longitudinal Freeurfer (SS=152) and Jacoby Integration (SS=138) to SS=125 subjects enrolled. Atrophiq release 2.0 using symmetric stationary velocity fields, Wendland kernel regularization, and differential bias correction reduces this further.
Prognostic Markers
The hippocampal volume is often used as a prognostic marker for transition from MCI to AD. Atrophiq offers the possibility to quantify the structural integrity of the hippocampus by texture analysis and thereby make significantly more precise prognosis. The area under the receiver-operator characteristic curve (AUC) separation of MCI that are one-year stable compared to converters to AD is 0.78 on ADNI and 0.83 on the AIBL data set. This persists after adjustment for all major risk factors (Age, gender, hippocampal volume and MMSE). Atrophiq prognosis makes it possible to maintain power while reducing cost by patient selection. Cost is reduced by 17% in a 2-year trial. A 1-year trial will have the same cost as a standard 2-year trial.
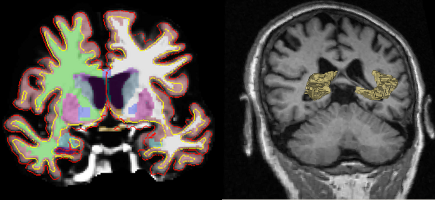
RIGHT: A MRI slice overlaid with a 3D rendering of the segmented hippocampus.